Table Of Content
We are not aware of any other general purpose public tool that has integrated similar functionality to design target-specific primers. A number of studies have investigated the effects of mismatches between targets and primers and have shown that a target can be amplified even if it has a few mismatches to the primers [2–5]. In general, mismatches towards the 3’ end affect target amplification much more than mismatches towards the 5’ end.
A Step-by-Step Guide to Designing qPCR Primers
Not surprisingly, part or all of the forward primers picked by Primer-BLAST are located in exon 2 and all reverse primers are on the junctions between exon 3 and 5 (since exon 4 is not present). The Polymerase Chain Reaction is a powerful tool in recombinant DNA technology. A double-stranded DNA molecule that contains the desired sequence is denatured. Oligonucleotide DNA primers are then hybridized to the single strands on either side of the sequence to amplified. Modify your flanking sequence design until it no longer inteferes with proper folding of ROI. You may also wish to design a second set of flanking sequences; measurements on the extra set will serve as a control for perturbations from the flanking regions.
Designing forward primer
The GAPDH gene can also be used as the target gene for the internal control 17,57. In order to identify regions of the viral genome, further analysis of the molecular targets is needed to achieve the highest possible diagnostic accuracy. The primer/probe set should be evaluated with further experiments, so that more sensitive and specific primers/probes can be developed. Ordering primers and probes to perform validation testing on functionality and potential contaminants is also recommended 73.

Primer concentration and quality
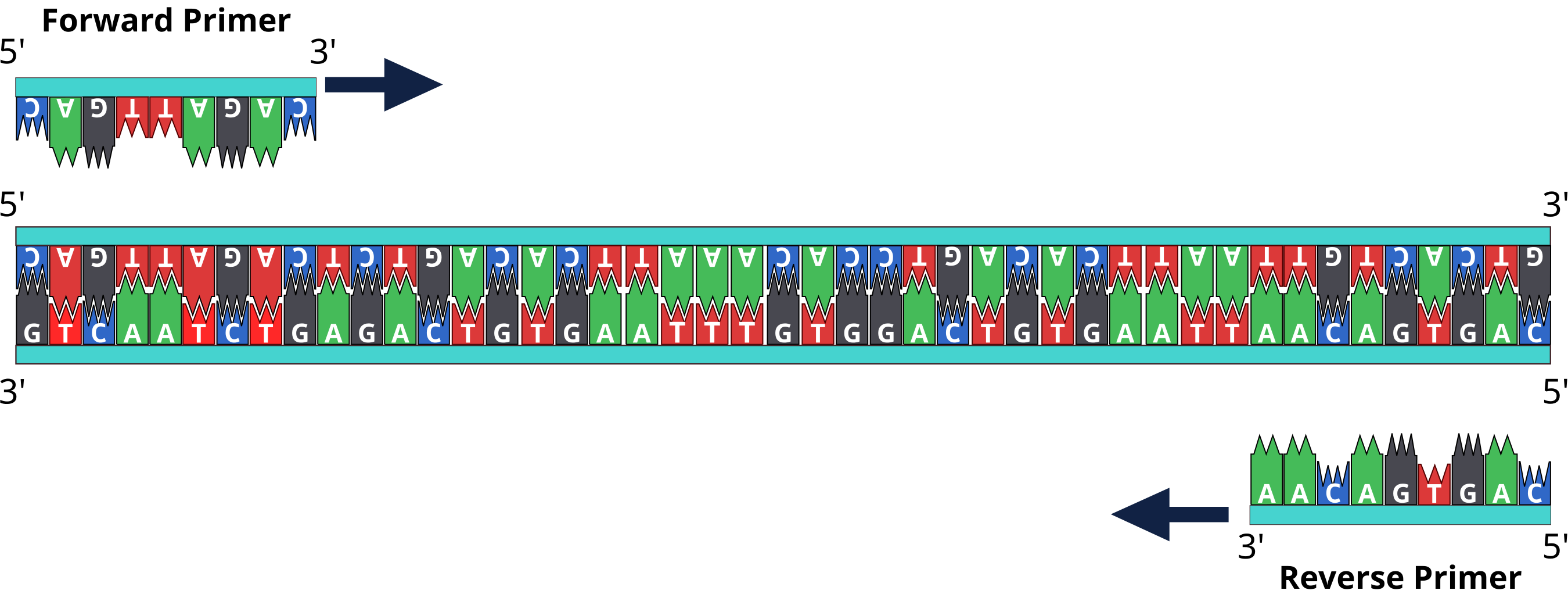
The 3′-OH ends of the primers face each other because they have annealed to opposite strands. The primers are then extended by DNA polymerase, creating two new double-stranded DNA molecules, each one containing a copy of the desired sequence or gene. Forward primers can also be called sense primers while the reverse primers are also known as the anti-sense primers. Primers generally bind to the opposite sides of the template DNA sequence or strands to be amplified during the PCR reaction. They are single-stranded (ss) DNA molecules or oligonucleotides, and they are synthesized by chemical processes.
To help read out the entire sequence of interest, and to save time on fluorescent labeling of primers, we add a 3´ tail extension with a shared primer binding site. This 3´ tail should be added to the DNA template sequence, and this will be transcribed into RNA. Before setting up the PCR experiment, the genomic DNA from both S. Cerevisiae and the Mycobacteriophage were quantified and diluted to a concentration that would allow between 104 and 107 molecules of DNA per reaction. A dilution to 10 ng/μl was generated by adding 48 μl into 452 μl of TE pH 8.0 buffer. Cerevisiae genome is about 12.5 Mb, 10 ng contain 7.41 X 105 molecules.
For longer templates, the primer range (upper right corner of Figure 1) should be used to limit the length. As aforementioned, primers are short DNA fragments (usually about bases or nucleotides long) that contain specific sequences which are complementary to the template DNA molecule to be copied and amplified. Two types of primers (i.e. the forward primers and the reverse primers) that are complementary to the 3′ end of each of the double stranded template DNA strands are used in any PCR reaction process. Generally, primers are short pieces of single-stranded DNA molecules that are usually around nucleotides in length.
The primer size for the forward and reverse primers can be specified (this size should not include the length of the overlap fragment). The 5′ and 3′ search areas are then specified by the user followed by the maximum and minimum percentage GC and primer Tm values for the forward and reverse primers. Primer Tm values are calculated using parameters defined previously7,8,9 and compare very closely with Tm values calculated by Primer310,11 (Fig. 2). Salt concentrations used during primer Tm calculations are set to 50 mM by default but can be changed to 10 mM.
Primer pairs should have melting temps (Tms) within 5°C of each other. These primer Tms influence the overall annealing temp (Ta) of the reaction. Our Tm calculator accounts for specific buffer compositions that impact primer melting temps. The goal is to avoid weak thermodynamics that can cause primer dimers to form due to inefficient binding. Enter the position ranges if you want the primers to be located on the specific sites. The positions refer to the base numbers on the plus strand of your template (i.e., the "From" position should always be smaller than the "To" position for a given primer).
An optimized protocol for stepwise optimization of real-time RT-PCR analysis Horticulture Research - Nature.com
An optimized protocol for stepwise optimization of real-time RT-PCR analysis Horticulture Research.
Posted: Sun, 01 Aug 2021 07:00:00 GMT [source]
Thus we conclude that Primer-BLAST is able to detect potential unintended targets that are missed by QuantPrime or PRIMERGENS during the specificity screening process. A different DNA template was used for the experiment presented in Figure 3b. Genomic DNA from a Mycobacteriophage was used to amplify a conserved 566 bp DNA segment. Like the previous experiment, the optimal Mg2+ concentration had to be determined. As shown in Figure 3b, amplification of the desired PCR product requires at least 2.0 mM Mg2+ (lane 5).
Has anyone made human RNAse P cDNA PCR primers? - ResearchGate
Has anyone made human RNAse P cDNA PCR primers?.
Posted: Wed, 02 Dec 2020 08:00:00 GMT [source]
There are several existing programs that have addressed some aspects of this issue. Autoprime [10] designs primers spanning exon junctions or introns so that the primers only target mRNA. QuantPrime [11] is a specialized tool to design target-specific primers for detecting mRNA in real time PCR. Likewise, the PRIMEGENS Sequence Specific Primer Design tool [12] can also be used for specific primer design for a limited number of organisms. Other limitations in these tools include low target detection sensitivity, limited specificity stringency options, no or limited support for designing primers based on exon/intron boundary requirements and limited coverage of organisms in search databases. Primer-BLAST is a general purpose target-specific PCR primer design tool that offers a level of sensitivity and usability not found in other tools.
This requires at least one primer (for a given primer pair) to have the specified number of mismatches to unintended targets. The larger the mismatches (especially those toward 3' end) are between primers and the unintended targets, the more specific the primer pair is to your template (i.e., it will be more difficult to anneal to unintended targets). However, specifying a larger mismatch value may make it more difficult to find such specific primers.
For example, if you want the PCR product to be located between position 100 and position 1000 on the template, you can set forward primer "From" to 100 and reverse primer "To" to 1000 (but leave the forward primer "To" and reverse primer "From" empty). Note that the position range of forward primer may not overlap with that of reverse primer. In addition to sensitivity, specificity is an important indicator when considering primers and probes. Whether there is cross-reactivity is an important aspect of evaluating specificity.
In case, if you take the complementary sequence and use it for primer designing (laying the sequence in 5’-3’), the reverse primer in the above example now acts as the forward primer. For structure mapping experiments, the modified RNA is reverse transcribed into DNA, which is then read by a sequencer. Reverse transcription requires the presence of a primer (18 – 20 bases), complimentary to the 3´ end of the RNA, which is then extended all the way to the 5´ end of the RNA, or to other stops within the RNA sequence, depending upon the modification being used. This means that modifications in the last ~20 bases in the RNA, cannot be read out using primer extension.
No comments:
Post a Comment